Protein Dynamics at Interfaces
Protein Dynamics at Interfaces
Interactions at surfaces define the reactivity of proteins, whether this be plastic wall in your milk jug, the polymer contact in your eye, or the permeable membrane in a chromatographic column. Our research attempts to characterize the dynamics of these interactions using single-molecule, super-resolution fluorescence microscopy. In doing so, we hope to identify the mechanisms that foul membranes, inhibit protein motion, and detract from successful separations. We can achieve this by considering the protein conformational changes, kinetics of protein interactions, and the motion of proteins exploring the surface.
Approach
- FRET studies examine the opening and closing of proteins in response to interactions with other substrates using Forster Resonance Energy Transfer (FRET).
- Kinetics of single-site adsorption events were monitored to elucidate the effects of changing the ionic content of the surrounding solution.
- Surface residence time distributions are monitored to investigate how functionalization of a chemical surface changes a proteins ability to explore a surface.
Related Papers
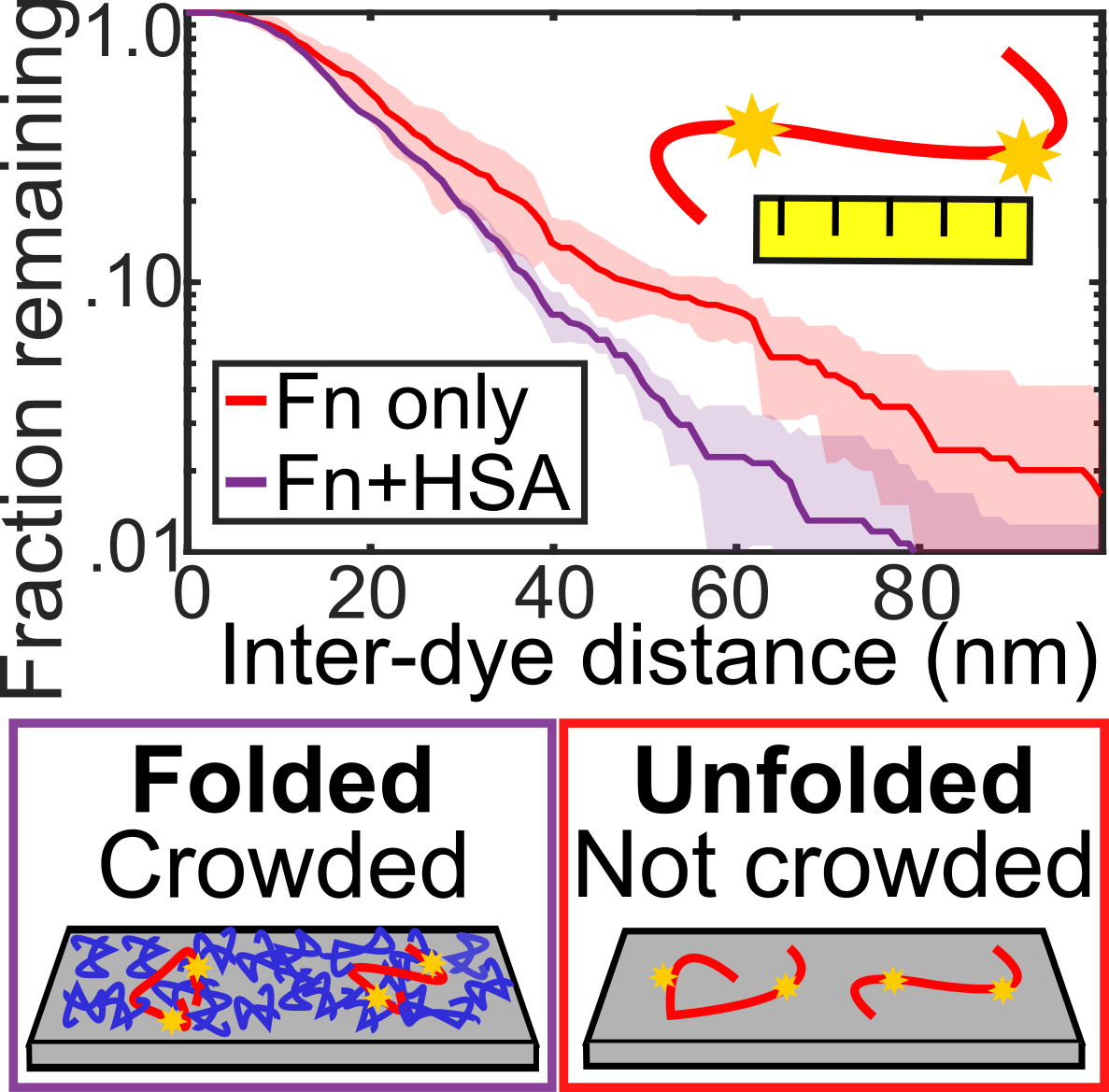
Warning, L.A., Zhang, Q., Baiyasi, R., Landes, C.F., Link, S., "Nanoscale Surface-induced Unfolding of Single Fibronectin is Restricted by Serum Albumin Crowding" JPCL 2020, 11, 3, 1170-1177.
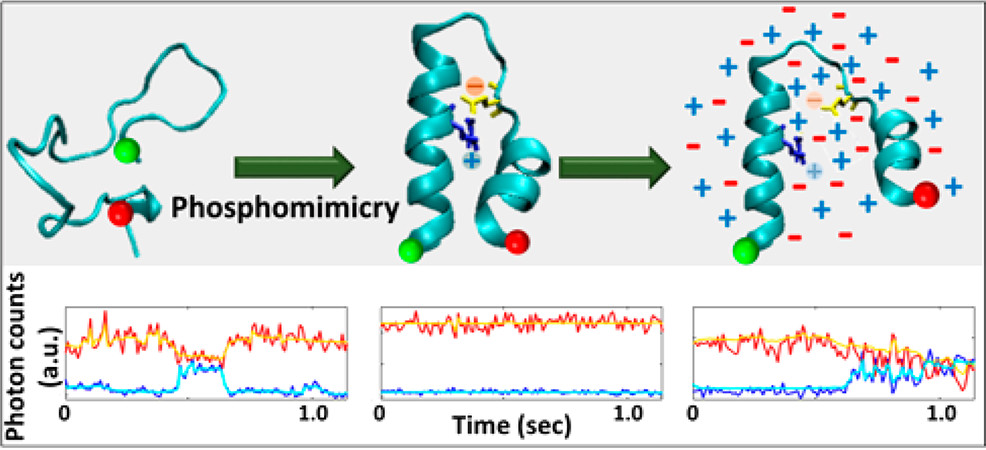
Chatterjee, S., A., Carina, Nurik, C.E., Carrejo, N.C., Dutta, C., Jayaraman, V., Landes, C.F., "Phosphorylation Induces Conformational Rigidity at the C-Terminal Domain of AMPA Receptors" J. Phys. Chem. B 2019
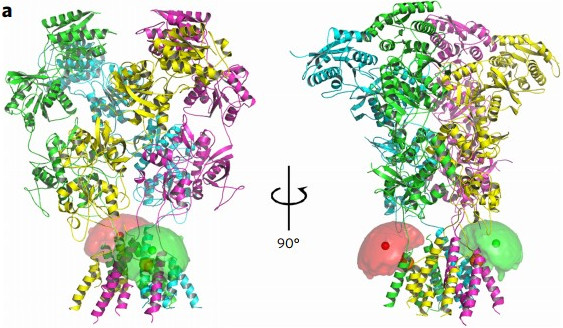
Dolino, D.M., Chatterjee, S., MacLean, D.M., Flatebo, C., Bishop, L.D.C., Shaikh, S.A., Landes, C.F., Jayaraman, V., "The structure–energy landscape of NMDA receptor gating" Nat. Chem. Bio., 2017, 13, pp. 1232-1238
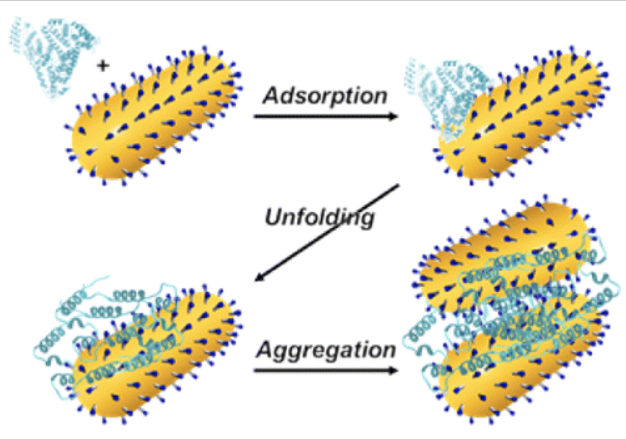
Dominguez-Medina, S.; Kisley, L.; Tauzin, L.J.; Hoggard, A.; Shuang, B.; Indrasekara, A.S.D.S.; Wang, L.-Y.; Chen, S.; Derry, P.J.; Liopo, A.; Zubarev, E.R.; Landes, C. F.; Link, S., "Adsorption and Unfolding of a Single Protein Triggers Nanoparticle Aggregation" ACS Nano 2016, 10, 2103-2112.
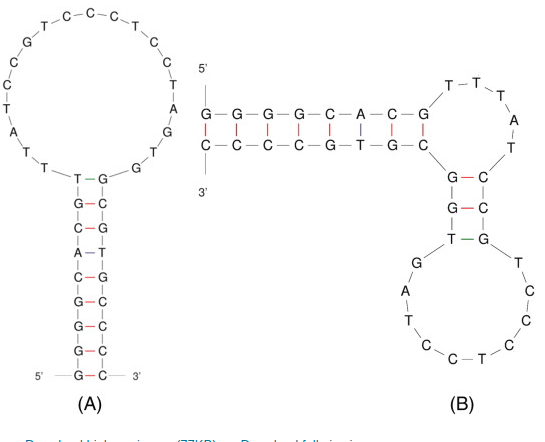
Poongavanam, M. -V.; Kisley, L.; Kourentzi, K.; Willson, R.C.; Landes, C. F., "Ensemble and single-molecule biophysical characterization of D17.4 DNA aptamer–IgE interactions" BBA Proteins Proteom. 2016, 1864, 154-164.